Introduction
Magen has been developing nucleic acid extraction technologies for over ten years, with a consistent focus on membrane-based purification systems. The HiPure Circulating DNA Kit (IVD3182) represents the column-based format within the circulating DNA portfolio, designed for plasma and serum workflows ranging from 1–5 mL input volumes.
The kit utilizes a vacuum-assisted silica membrane purification system optimized for recovery of low-abundance circulating DNA fragments while maintaining stable filtration performance. Membrane architecture and binding buffer formulation have been refined through internal optimization and validation experiments.
In internal verification testing, short DNA fragment recovery efficiencies typically exceeded 80% under controlled extraction conditions. Plasma extraction experiments using 5 mL input volumes showed stable cfDNA concentrations across replicate samples, with performance comparable to established column-based circulating DNA purification systems processed under equivalent conditions.
· Laboratories operating different workflow formats may refer to:
· Smaller plasma volumes 0.6mL→ ( D3180 )
· 1mL processing → ( D3181 )
· Centrifuge-based workflow → ( D3182D )
· Magnetic bead automation → ( IVD5435 )
Details
Workflow
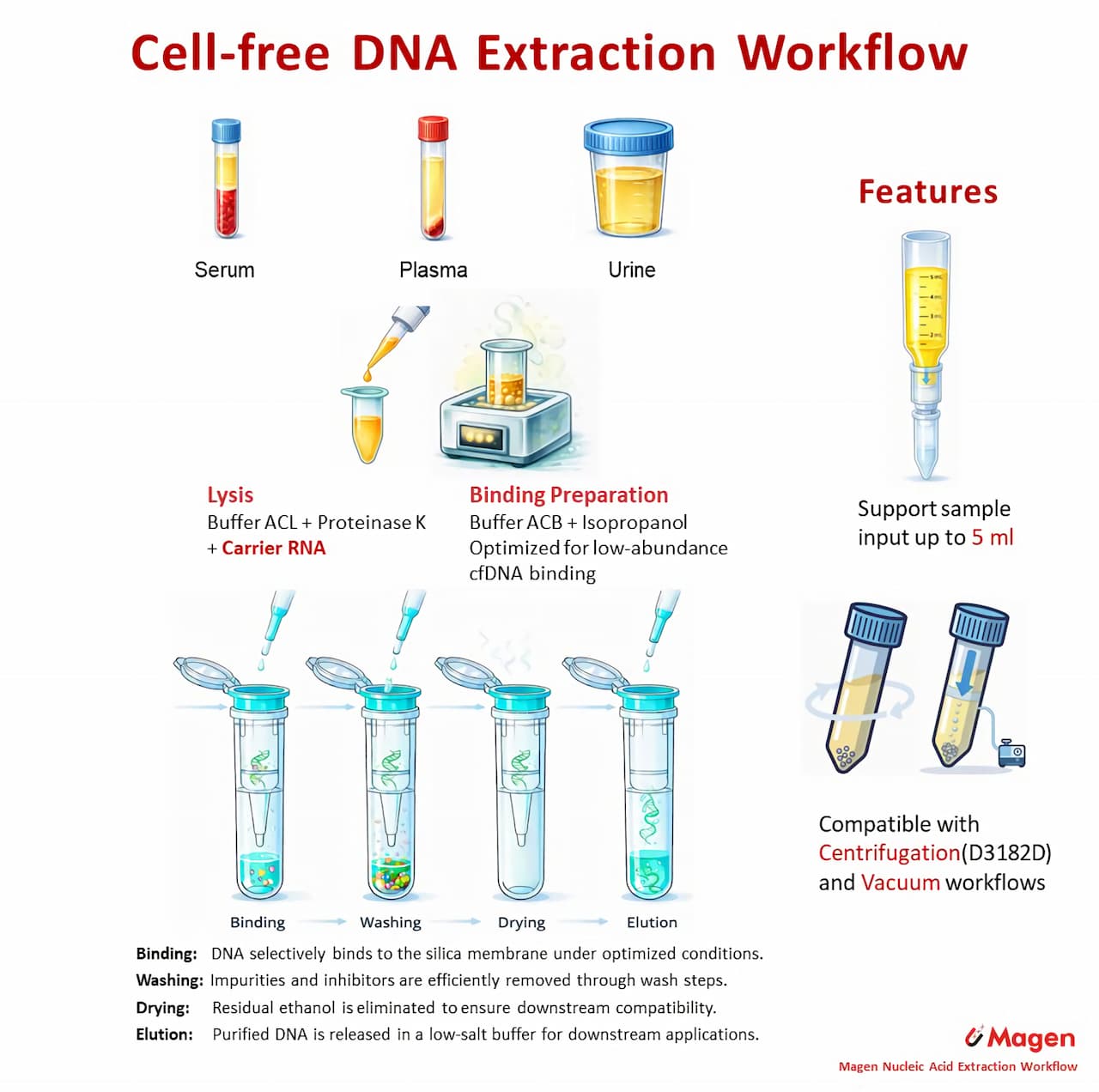
Workflow Overview
The HiPure Circulating DNA Kit uses a silica column–based workflow for purification of circulating DNA from plasma and serum samples. Following carrier RNA–assisted lysis, DNA is captured on the silica membrane under optimized binding conditions and recovered through washing, drying and elution steps. This workflow provides a structured and reproducible approach for routine cfDNA extraction. A centrifugation-based variant is available as D3182D for laboratories that prefer spin-based handling over vacuum processing.
Sample Handling Logic
The workflow is designed for 1–5 mL plasma or serum input, with the main variation concentrated in the lysis and binding stages under larger-volume handling. Once DNA binding conditions are established, the downstream column purification steps remain consistent, enabling stable performance across routine sample conditions.
Time and Workflow Characteristics
Under typical manual operation, the overall workflow is usually completed within about 65–80 minutes, depending mainly on sample input volume and lysis conditions. For detailed step-by-step conditions, workflow guidance and estimated processing times, please refer to the Workflow Note in the Download section.
Specifications
|
Features
|
Specifications
|
|
Main Functions
|
Isolation circulating DNA from 1-5ml plasma, serum, body fluids using vacuum protocol
|
|
Applications
|
qPCR, liquid or solid chip analysis, hybridization and SNP detection, etc.
|
|
Purification method
|
Mini spin column
|
|
Purification technology
|
Silica technology
|
|
Process method
|
Manual (vacuum)
|
|
Sample type
|
Serum, plasma and other cell-free fluid samples
|
|
Sample amount
|
1-5ml
|
|
Elution volume
|
≥50μl
|
|
Time per run
|
65-80 minutes
|
|
Liquid carrying volume per column
|
4ml
|
|
Binding yield of column
|
1mg
|
Extraction Principle
The HiPure Circulating DNA Kit is designed for purification of circulating cell-free DNA (cfDNA) from plasma, serum, or urine samples. Circulating nucleic acids in biological fluids are typically present at low concentrations and often exist as short DNA fragments.
Samples are lysed using a chaotropic buffer together with Proteinase K to release nucleic acids from protein complexes and protect DNA from degradation. Carrier RNA is included to improve recovery efficiency of low-abundance DNA molecules.
Under optimized binding conditions, circulating DNA fragments bind to the silica membrane of the HiPure cfDNA column while proteins and other contaminants pass through during filtration. After sequential washing steps to remove residual impurities, purified DNA is eluted in a low-salt buffer suitable for downstream molecular analysis workflows.
Engineering Characteristics
Optimized Column Membrane Architecture
Internal evaluation of different membrane configurations showed that a three-layer silica membrane structure provided an effective balance between DNA adsorption efficiency and filtration time during cfDNA extraction.
Binding Buffer Optimization
An updated binding buffer formulation (ACB) was developed to reduce solution viscosity during plasma processing. Internal testing showed that this modification improved filtration efficiency while maintaining comparable DNA recovery performance.
Designed for Large-Volume Plasma Processing
The column format supports processing of milliliter-scale plasma samples, enabling recovery of low-abundance circulating DNA molecules commonly present in clinical liquid biopsy samples.
System Integration
IVD3182 connects to a structured circulating DNA product framework:
-
Low-volume plasma extraction → (D3180)
-
Standard 1mL workflow → (D3181)
-
Centrifuge processing → (D3182D)
-
Scalable magnetic workflow → (IVD5435)
-
Fragment selection workflows → (12927)
Technical Validation
Validation experiments were performed to evaluate recovery efficiency and reproducibility of the purification system.
Short DNA Recovery Testing
Marker recovery experiments demonstrated DNA recovery efficiencies generally exceeding 80%, confirming effective capture of fragmented DNA molecules typical of circulating nucleic acids.
Batch Consistency Testing
Batch-to-batch verification experiments showed stable DNA recovery performance across different production lots, with recovery rates consistently above 80% and minimal variation between tested batches.
Urine Sample Validation
Urine extraction experiments using spiked short DNA fragments demonstrated recovery efficiencies typically ranging from 84% to 102% under optimized extraction conditions.
Customer Sample Validation
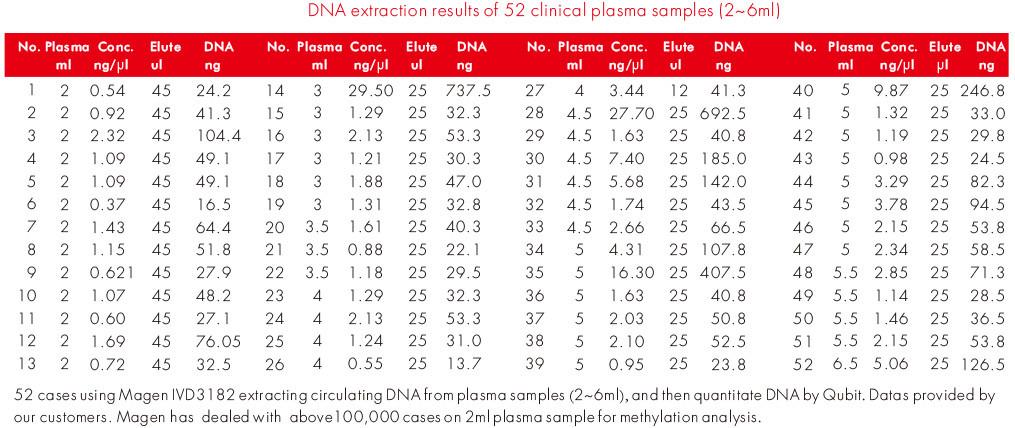
Performance was further evaluated using 46 customer plasma samples processed with the HiPure Circulating DNA Kit under routine testing conditions at a leading oncology testing company. Among these samples, 45 were assessed as qualified and 1 as risk, indicating stable extraction performance across real-world sample inputs.
Sample input volumes ranged from 3.0 mL to 6.5 mL, with most workflows centered around 4.5-5.0 mL plasma. These results support the suitability of the system for circulating DNA extraction from clinically relevant plasma volumes.
Fragment Size Recovery
Circulating cell-free DNA molecules are typically fragmented and often fall within the nucleosomal DNA size range.
Fragment recovery experiments using short DNA markers confirmed that the purification system can capture fragments around 50 bp in length with recovery efficiencies above 80%.
Analysis of circulating DNA extracted from serum samples showed fragment distributions typically ranging between approximately 150 bp and 1000 bp, consistent with reported cfDNA fragment characteristics.
These results indicate that the purification chemistry supports recovery of fragmented circulating DNA suitable for downstream molecular analysis.
Performance Comparison
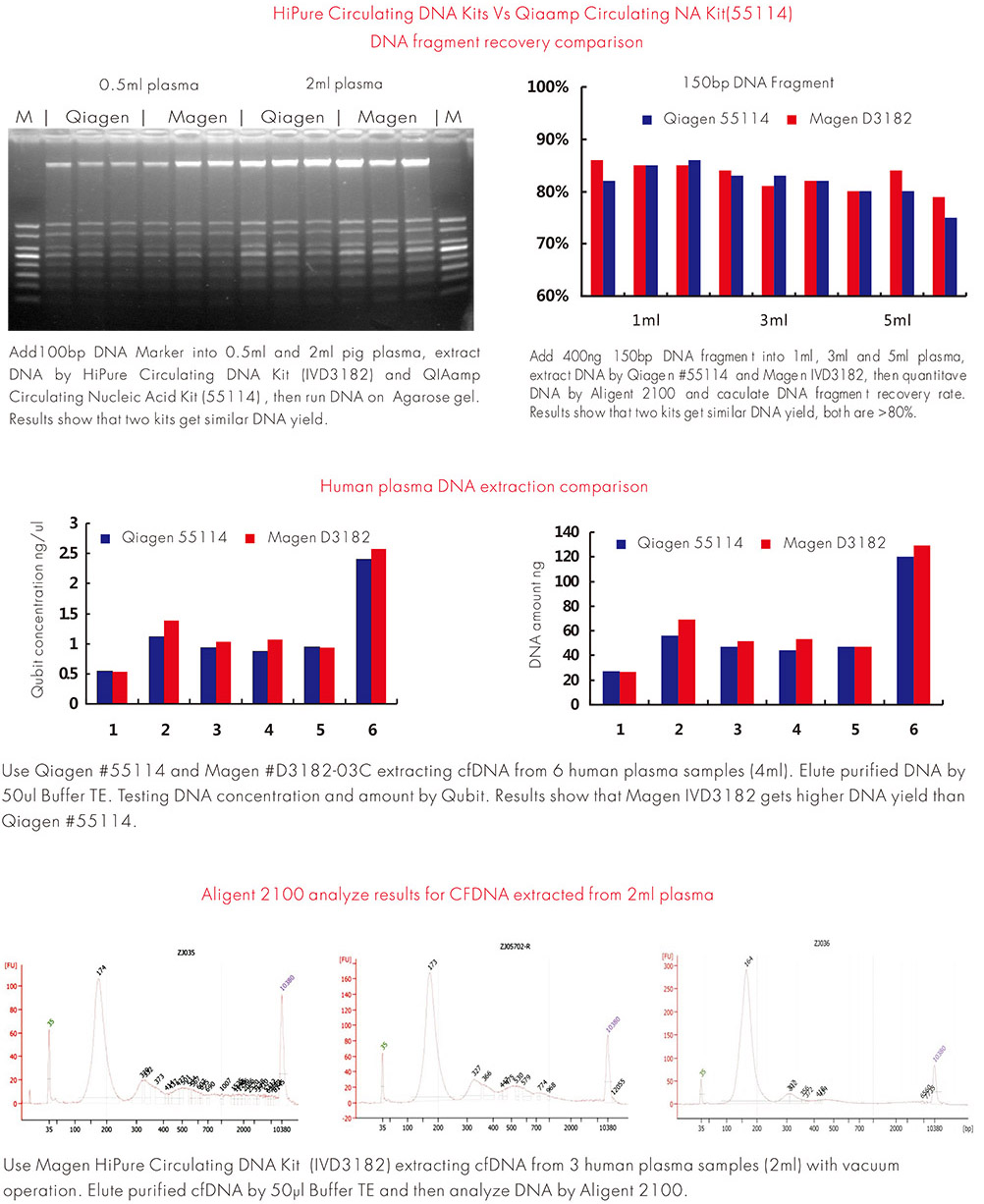
Comparative extraction experiments were performed using plasma samples processed with the HiPure Circulating DNA Kit and a widely used commercial cfDNA extraction system.
In testing using 5 mL plasma samples, Qubit quantification indicated that DNA concentrations obtained with the HiPure system were comparable to or higher than those obtained with the reference column-based extraction kit in several replicate samples.
Downstream library preparation experiments demonstrated that extracted DNA supported adapter ligation and PCR amplification workflows used for sequencing library construction.
Application Scenario Summary
The following table summarizes selected published research scenarios in which IVD3182 or closely related HiPure circulating DNA formats were used for cfDNA / ctDNA preparation. The purpose is to help users understand typical application directions of this product route before reviewing the related technical notes and publications.
|
Application Scenario
|
Related Format
|
Sample / cfDNA Source
|
Downstream Research Use
|
|
Advanced NSCLC ctDNA profiling for brain metastasis–associated genomic alteration and tissue–plasma concordance research
|
D3182C
|
Plasma-derived ctDNA from advanced NSCLC patients with or without brain metastases
|
95-gene targeted NGS for somatic mutation profiling, brain metastasis–associated genomic alteration comparison, tissue–plasma concordance analysis and actionable alteration assessment
|
|
NSCLC ctDNA and tissue DNA comparison for actionable genomic alteration profiling and EGFR-TKI response research
|
D3182C
|
Plasma-derived ctDNA from NSCLC patients, compared with tumor tissue DNA
|
95-gene targeted NGS for tissue–plasma mutation concordance analysis, LUAD/LUSC subtype comparison, actionable alteration assessment and EGFR-TKI response association research
|
View more application scenarios
|
Application Scenario
|
Related Format
|
Sample / cfDNA Source
|
Downstream Research Use
|
|
ALK-positive advanced NSCLC ctDNA dynamic monitoring for lorlatinib treatment response and acquired resistance profiling
|
D3182
|
Serial plasma ctDNA from ALK-positive advanced NSCLC patients during lorlatinib treatment
|
556-gene targeted NGS for serial ctDNA VAF monitoring, lorlatinib response assessment and acquired ALK resistance mutation profiling
|
|
AITL cfDNA liquid biopsy for mutation profiling, prognosis stratification and potential PD-1 response biomarker research
|
D3182
|
Pretreatment peripheral blood cfDNA from newly diagnosed AITL patients, with paired tumor tissue DNA
|
Customized 46-gene targeted NGS for cfDNA–tissue concordance analysis, recurrent mutation discovery and prognosis / PD-1 response biomarker research
|
|
Postoperative ctDNA recurrence prediction in CRLM patients after hepatectomy
|
D3182D
|
cfDNA / ctDNA from 3–5 mL postoperative plasma of CRLM patients
|
Dual-index UMI ultra-deep targeted NGS with a CRLM-specific J25 panel for postoperative ctDNA detection, recurrence prediction and DFS / OS stratification
|
|
EOC plasma cfDNA methylation marker discovery for early ovarian cancer detection model development
|
D3182
|
Plasma cfDNA from EOC patients and healthy female controls
|
cfDNA methylation marker screening, MethylBERT-based EOC diagnostic model development, prognostic analysis and OV1 ddPCR assay adaptation
|
|
CRC MYO1-G ctDNA methylation marker validation for diagnosis, tumor burden assessment and treatment-response monitoring
|
D3182
|
Plasma cfDNA from CRC patients and tumor-free controls
|
ddPCR-based MYO1-G methylation assay for CRC diagnosis, tumor burden assessment, post-surgery monitoring and treatment-response evaluation
|
|
MCED-oriented cfDNA cleavage-profile analysis for WGS-based methylation-status prediction
|
D3182S
|
cfDNA from 3 mL plasma of healthy individuals and patients with HCC, lung cancer or colorectal cancer
|
WGS / WGBS comparison and XGBoost modeling to predict individual CpG methylation status from cfDNA cleavage profiles for MCED biomarker development
|
|
HBV-related HCC detection and monitoring using plasma cfDNA fragmentation profiles
|
D3182S
|
cfDNA from 3 mL plasma of healthy individuals, hepatitis B patients, cirrhosis patients and HBV-related HCC patients
|
Low-coverage WGS and machine-learning analysis of cfDNA fragmentation profiles for HBV-related HCC detection, disease-stage profiling and relapse-related assessment
|
|
Maternal plasma cfDNA allele fraction analysis for single-gene disorder NIPT accuracy improvement
|
D3182
|
Maternal plasma cfDNA from pregnant women at risk of fetal monogenic disorders
|
Unique dual indexing targeted deep sequencing to reduce chimeric amplicon contamination and improve fetal allele fraction measurement for TSC / Marfan syndrome inheritance and de novo mutation analysis
|
|
Maternal plasma evDNA and cfDNA comparison for prenatal genetic disease proof-of-concept research
|
D3180
|
evDNA from maternal plasma EVs and matched plasma cfDNA from 250 μL maternal plasma
|
qPCR, paired-end low-coverage WGS and targeted FGFR3 amplicon sequencing to compare evDNA with cfDNA and evaluate proof-of-concept detection of fetal sex, trisomy 21/18/13 and de novo FGFR3 mutations
|
Note: These examples describe published research uses of IVD3182 or closely related HiPure circulating DNA formats for upstream cfDNA / ctDNA preparation. The extraction workflow provides purified nucleic acid input for downstream assays; final research conclusions depend on the complete study design, sequencing or detection method, bioinformatics analysis and clinical validation strategy.
Kit Contents
|
Contents
|
IVD3182
|
|
Purification Times
|
50
|
|
Buffer ACL
|
250 ml
|
|
Buffer ACB*
|
300 ml
|
|
Buffer DCW1*
|
22 ml
|
|
Buffer DCW2*
|
10 ml
|
|
Proteinase K
|
540 mg
|
|
Protease Dissolve Buffer
|
30 ml
|
|
Carrier RNA
|
110 μg
|
Nuclease Free Water
|
20 ml
|
HiPure CFDNA Mini Columns
|
50
|
2 ml Collection Tubes
|
100
|
Extender Tube
|
50
|
Vac-Connector
|
50
|
Storage and Stability
Proteinase K, Carrier RNA should be stored at 2-8°C upon arrival. However, short-term storage (up to 12 weeks) at room temperature (15-25°C) does not affect their performance. The remaining kit components can be stored dry at room temperature (15-25°C) and are stable for at least 18 months under these conditions. The entire kit can be stored at 2-8°C, but in this case buffers should be redissolved before use. Make sure that all buffers are at room temperature when used.
Experiment Data



.jpg)


