1.0 Introduction
Circulating cell-free DNA workflow design is shaped less by sample type than by a small group of technical variables: plasma input volume, target DNA abundance, fragment profile and downstream analytical requirements. These variables determine how DNA should be captured, how much sample can be handled in a practical workflow, and whether standard recovery or fragment enrichment is the more appropriate route.
Within the Magen cfDNA system, three technical routes are used: column-based extraction, magnetic bead-based extraction and fragment enrichment. These routes are not simply different kit formats. They represent different technical solutions to different cfDNA handling problems.
2.0 Technical Architecture of the Magen cfDNA Workflow System
The Magen cfDNA system can be understood as three related but distinct workflow routes.
2.1 Column-based extraction
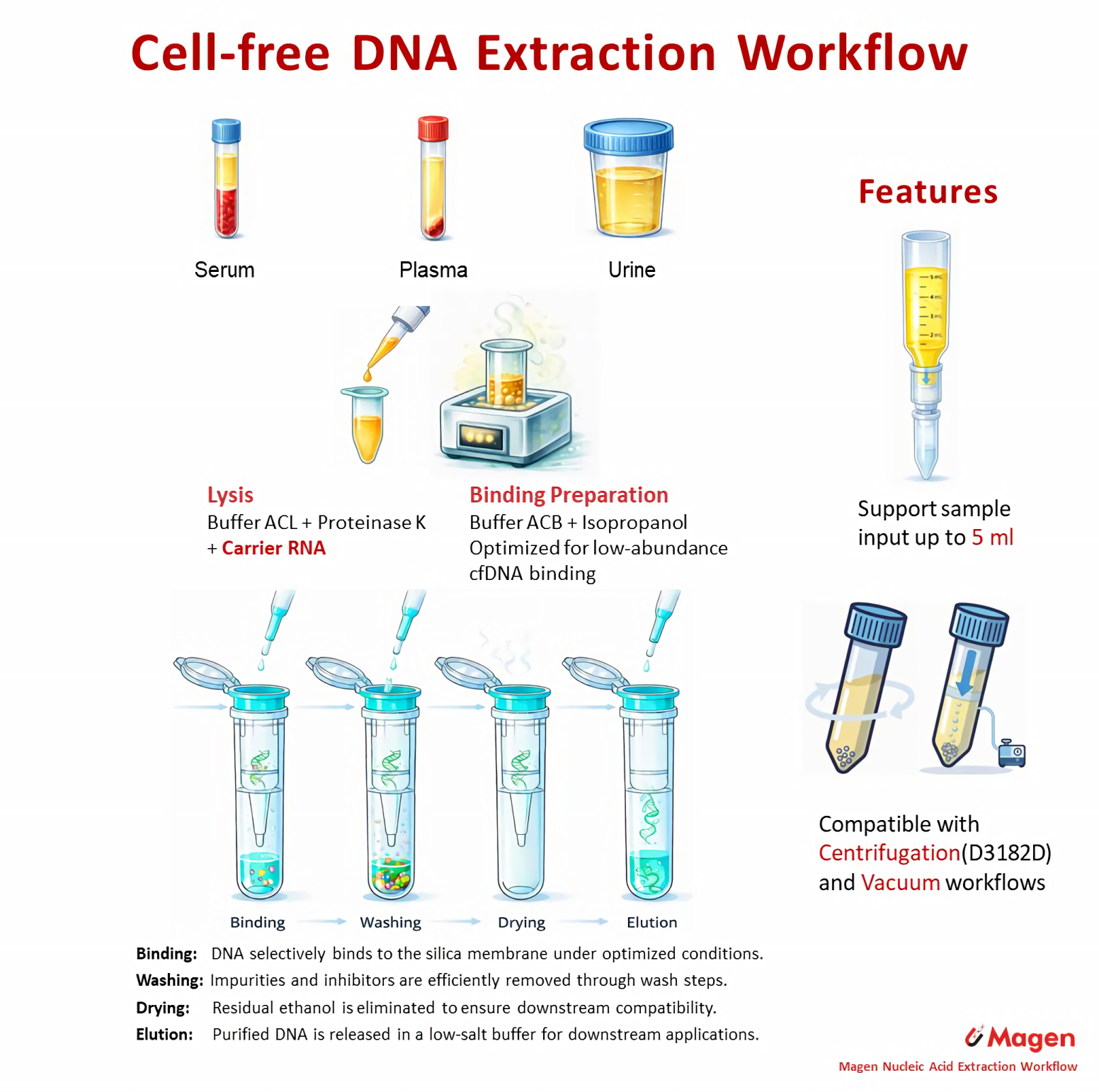
Column-based workflows use silica membrane adsorption under flow-through conditions. DNA binding occurs as the lysate passes through the membrane, which gives this route a structured and reproducible handling pattern. Within the Magen system, this route covers the 1–5 mL large-volume core workflow together with smaller-input column branches.
Representative workflow: HiPure Circulating DNA Kit (IVD3182)
2.2 Magnetic bead-based extraction
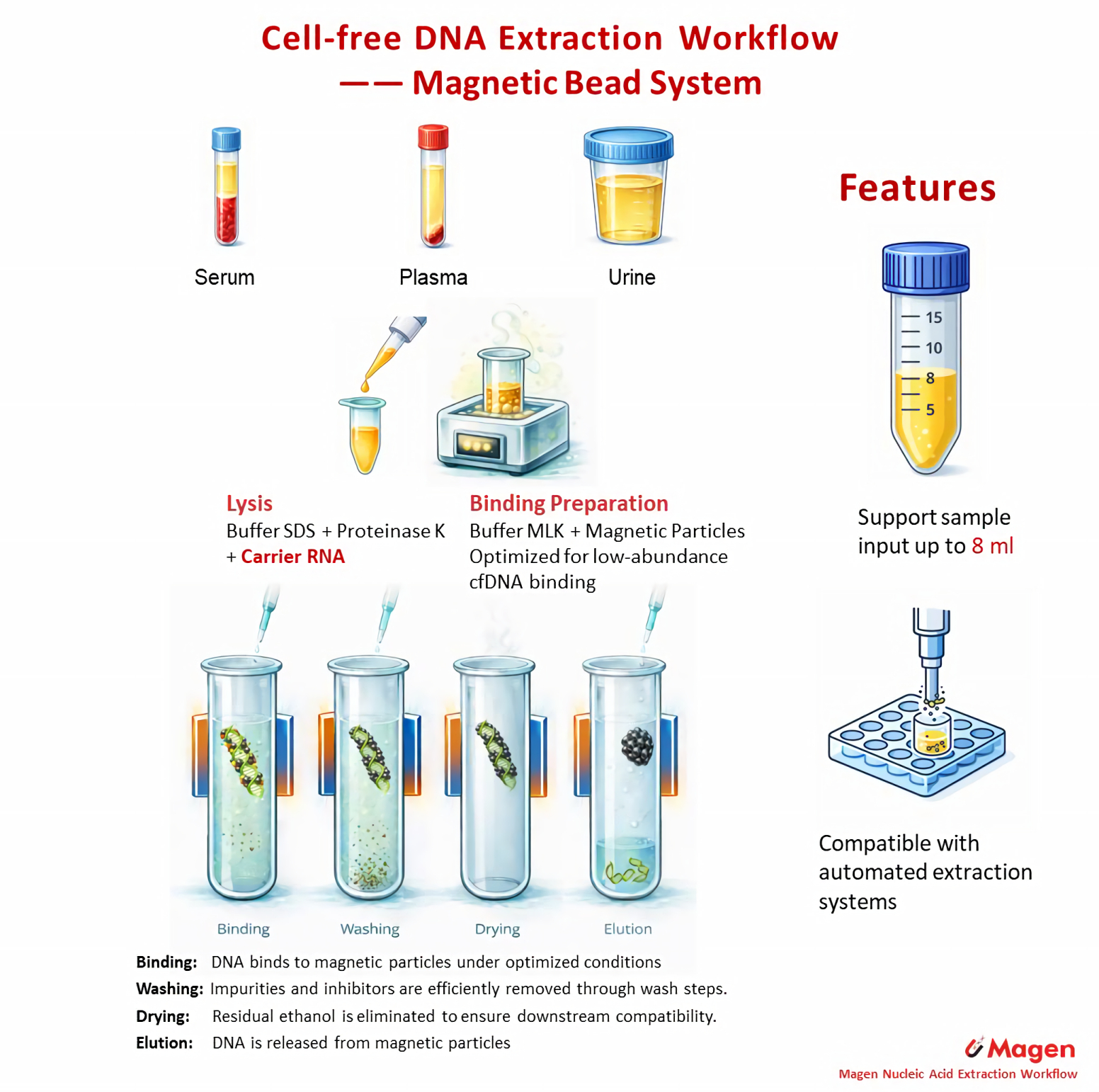
Magnetic bead-based workflows rely on particle–DNA interaction under mixing conditions rather than membrane flow-through. This allows more flexible scaling for larger plasma volumes and provides a practical route for low-abundance cfDNA recovery. Within the Magen system, this route includes both large-volume and low-input magnetic workflows.
Representative workflow: MagPure Circulating DNA Maxi Kit (IVD5435)
2.3 Fragment enrichment
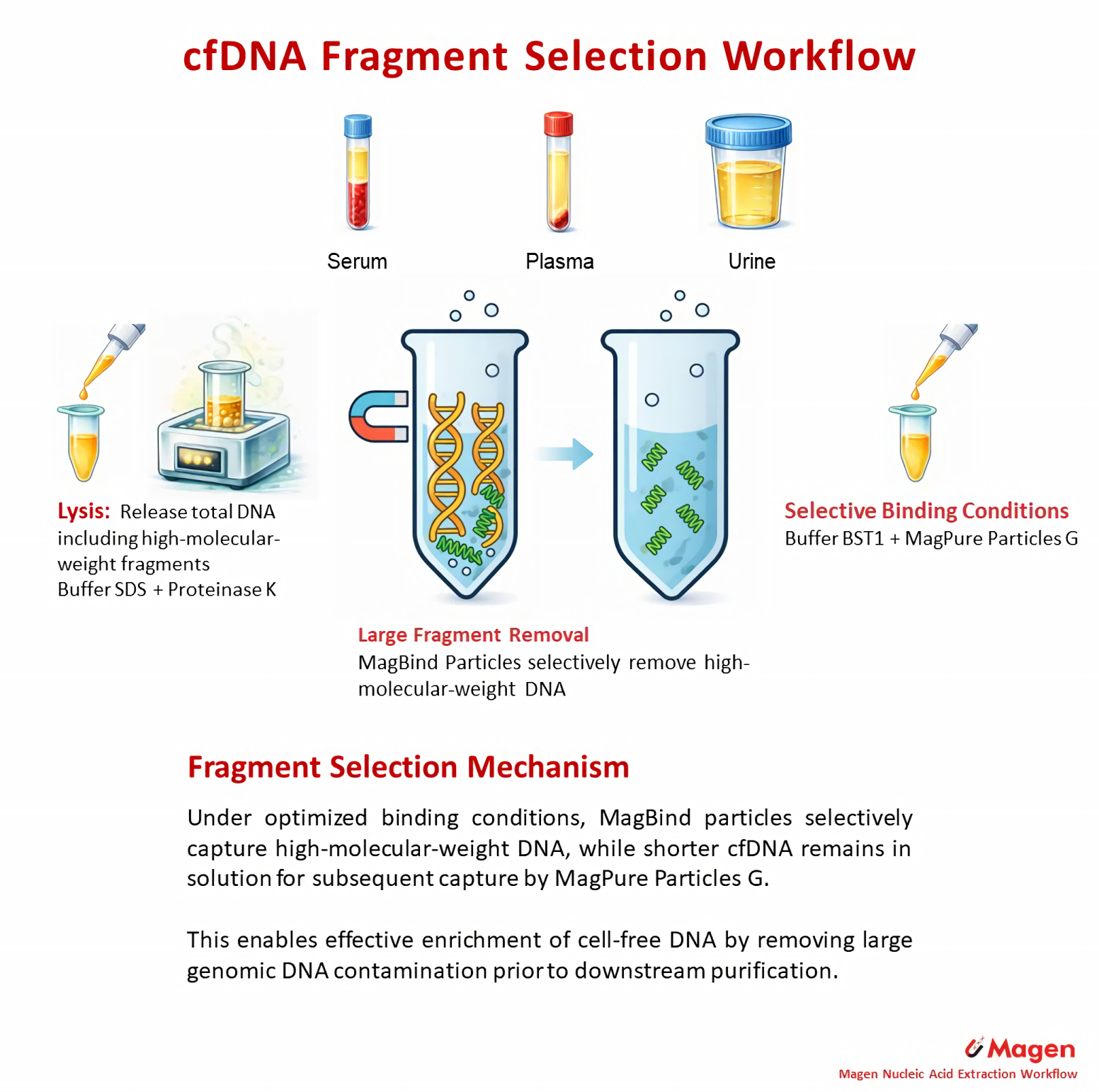
Fragment enrichment workflows add a size-selection step before final capture. Instead of recovering total cfDNA directly, they first reduce high-molecular-weight DNA background and then enrich shorter cfDNA fractions. This makes them a distinct route rather than a simple extension of standard extraction.
Representative workflow: MagPure Circulating DNA Rich Maxi Kit (12927)
3.0 Workflow Development Background
The current cfDNA workflow system did not arise from a single extraction logic applied across all conditions. It took shape because different problems started to matter once input volume increased, DNA abundance dropped, or fragment profile itself became part of the analytical target.
In the column-based route, the question was never only whether cfDNA could bind to a silica membrane. The harder question was how stable that capture would remain once the workflow moved into larger-volume handling. This is where details that look small on paper started to matter: membrane behavior during prolonged flow-through, differences between column structures, and how Buffer ACB and ACB2 changed the balance between filtration speed and recovery consistency. Once plasma volume increased, speed and recovery no longer moved together automatically, so the route had to be tuned around both.
The magnetic bead route developed under a different pressure. Here the issue was not membrane passage, but how far particle-based capture could be extended before the workflow itself needed to branch. At lower input, a single binding step was usually enough. At higher plasma volumes, the same logic no longer remained equally robust in practice, and staged binding became part of the route design. From there, other details became difficult to ignore, including particle response, reagent stability, and whether batch behavior remained consistent over time. These are not peripheral points in the magnetic route. They are part of what makes it workable as a scalable system.
The enrichment route came out of yet another practical question. Once fragment profile became something to control rather than simply accept, total recovery alone stopped being a sufficient endpoint. What started to matter was how far large-fragment depletion could be pushed, how Select Solution level changed cutoff behavior, and how much short-fragment recovery could still be preserved after selection. That is why staged particle handling and cutoff tuning sit near the center of this route. Without these elements, enrichment would be difficult to distinguish from a standard magnetic workflow. With them, it becomes a separate technical branch built around fragment behavior itself.
Taken together, these routes reflect not just different product formats, but different technical answers to different cfDNA handling problems. That is why the current Magen cfDNA portfolio is organized as a workflow system rather than a single extraction format with minor variations.
4.0 Representative Workflows in the Magen System
4.1 Column-based core workflow
HiPure Circulating DNA Kit (IVD3182) represents the core large-volume column route in the Magen cfDNA system. It supports 1–5 mL plasma or serum input and uses carrier RNA-assisted lysis followed by silica membrane capture. A centrifuge-based variant is available through D3182D, while D3181 and D3180 extend the same general column logic to routine ~1 mL and lower-input ~0.6 mL workflows.
4.2 Magnetic bead-based core workflow
MagPure Circulating DNA Maxi Kit (IVD5435) represents the core magnetic bead route. It supports 1–8 mL input and uses extended bead interaction rather than membrane flow-through. For 1–4 mL input, binding is typically completed in a single step. For volumes above 4 mL, the workflow branches into two sequential binding steps. The lower-input magnetic branch is represented by IVD5432.
4.3 Fragment enrichment core workflow
MagPure Circulating DNA Rich Maxi Kit (12927) represents the enrichment route. Its defining feature is not simply magnetic purification, but sequential size-selective processing. MagBind particles are first used to reduce larger DNA fragments, after which shorter cfDNA fractions are captured with MagPure particles. The lower-input branch of this route is represented by 12917.
5.0 Workflow Time Comparison
Typical processing time differences arise from the number of incubation blocks, the binding strategy and the complexity of downstream handling.
|
Workflow Route |
Representative Product |
Representative Input |
Typical Processing Time |
|
Column-based extraction |
IVD3182 |
5 mL |
~65–80 min |
|
Magnetic bead-based extraction |
IVD5435 |
5 mL |
~105–130 min |
|
Fragment enrichment |
12927 |
5 mL |
~110–140 min |
Column workflows are usually faster because DNA capture occurs during membrane passage. Magnetic workflows require more handling time because binding depends on mixing, magnetic separation and repeated liquid handling. Enrichment workflows add a size-selection stage before final capture, so they are typically the most time-intensive route.
Detailed Workflow Notes
The representative routes outlined above are also available as detailed workflow notes. These pages provide step-by-step handling logic, estimated processing time and route-specific workflow structure for the three core cfDNA workflows in the Magen system.
HiPure Circulating DNA Kit (IVD3182) — detailed column-based workflow
MagPure Circulating DNA Maxi Kit (IVD5435) — detailed magnetic bead-based workflow
MagPure Circulating DNA Rich Maxi Kit (12927) — detailed fragment enrichment workflow
6.0 Pre-analytical and Workflow Considerations
Pre-analytical handling has a direct effect on cfDNA yield, fragment distribution and background genomic DNA contamination. Key variables include anticoagulant type, time from collection to plasma separation, freeze–thaw history and control of cellular lysis. These factors do not change the workflow route itself, but they can strongly influence recovery quality and downstream analytical sensitivity.
Common workflow risks are also route-dependent. In downstream processing, insufficient washing or drying may reduce assay compatibility, while incomplete mixing or magnetic separation can affect recovery consistency in bead-based workflows. For fragment-enrichment workflows, stronger size selection may improve short-fragment representation while reducing recovery of longer cfDNA fractions. For this reason, cfDNA workflow performance should be interpreted not only by total yield, but also by background control, fragment behavior and downstream fit.
7.0 Downstream Application Orientation
The three routes are not interchangeable in every context. Their technical differences create different strengths in downstream use.
| Workflow Route | Typical Downstream Fit | Why This Route Fits |
|---|---|---|
| Column-based extraction | Routine PCR, qPCR, general cfDNA preparation | Structured handling, reproducible processing and efficient routine recovery |
| Magnetic bead-based extraction | Large-volume plasma processing, low-abundance cfDNA recovery, automation-oriented workflows | Flexible scaling and extended particle interaction improve handling of larger inputs |
| Fragment enrichment | NGS, liquid biopsy, ctDNA-focused analysis, short-fragment-biased applications | Large-fragment depletion improves representation of short cfDNA fractions and reduces background |
These application mappings should be understood as workflow tendencies rather than absolute boundaries. Actual route selection still depends on sample condition, analytical sensitivity requirements and the balance between throughput and fragment control.
8.0 Closing Note
Taken together, these routes define a practical cfDNA workflow system spanning routine extraction, scalable large-volume recovery and fragment-oriented enrichment. Understanding where they diverge in capture logic, input handling and fragment behavior helps relate each route more clearly to the demands of downstream analysis.
Related cfDNA Application Resources
The appropriate cfDNA workflow depends on the downstream application. Column-based extraction, magnetic bead extraction and fragment-selective enrichment may serve different purposes in ctDNA mutation profiling, methylation biomarker research, fragmentomics, prenatal cfDNA analysis and emerging plasma cfDNA studies.
-
Real-World cfDNA Application Map
— overview of how cfDNA extraction requirements change across major research applications. -
ctDNA Extraction in Oncology Liquid Biopsy Research
— for mutation profiling, therapy resistance and MRD-related workflows. -
cfDNA Methylation Biomarkers in Cancer Research
— for methylation-sensitive plasma DNA workflows. -
cfDNA Fragmentomics and MCED Research
— for workflows where fragment size and cleavage patterns become part of the analytical signal. -
Maternal Plasma cfDNA Extraction for NIPS and NIPD Research
— for fetal cfDNA, fetal fraction and allele fraction workflows. -
Emerging Plasma cfDNA Research Beyond Cancer and NIPT
— for pregnancy complication, pathogen-derived and parasite-derived cfDNA research.


