FFPE Nucleic Acid Extraction Workflows Explained
Technical Route Design Behind the Magen FFPE Workflow System
1.0 Introduction
FFPE extraction is not a single protocol problem. It is a route-design problem shaped by fixation damage, analyte stability, partition strategy and downstream assay tolerance.
A paraffin block contains more than a target molecule. It contains a history of fixation, embedding, storage and tissue handling. Before extraction begins, nucleic acids may already be fragmented, crosslinked, chemically modified or associated with material that can affect binding, washing, quantification and amplification. A useful FFPE workflow therefore has to address the condition of the sample, not only the name of the sample type.
Within the Magen FFPE system, the workflow routes are not simple format variations. They represent different technical answers to different FFPE handling problems: routine DNA recovery, RNA recovery under controlled heat exposure, same-sample DNA/RNA partitioning, sequential magnetic adsorption, fragment-size selective DNA purification, high-purity DNA recovery and DNase-integrated RNA preparation.
The practical value of the system is found at the divergence points: where the sample is split, where binding occurs, how heat is used, how residual DNA or RNA is controlled, and how the purified nucleic acid is expected to behave in downstream analysis.
2.0 Technical Architecture of the Magen FFPE Workflow System
This route-design logic becomes clearer when the FFPE system is viewed through three major route groups: DNA-only workflows, RNA-only workflows and DNA/RNA co-extraction workflows. The same FFPE block can be used for DNA analysis, RNA analysis or combined DNA/RNA recovery, but these objectives do not place the same pressure on the extraction process.
DNA workflows usually need sufficient digestion and heat-assisted reversal of formalin-related modification. RNA workflows require controlled heat exposure and careful management of degradation risk. Co-extraction workflows must additionally decide where DNA and RNA should be separated within the process.
2.1 DNA-Only Workflow Routes
The DNA-only routes begin with a common FFPE challenge: paraffin must be removed, tissue must be digested, and formalin-related modification must be reversed sufficiently to recover amplifiable DNA. After this shared logic, the routes diverge according to the intended technical pressure.
The routine FFPE DNA column route is represented by D3126 HiPure FFPE DNA Kit. This route follows a classical silica membrane workflow after FFPE lysis and crosslink reversal. It is a linear DNA recovery route, where the main workflow concern is reliable DNA release, binding, washing and elution.
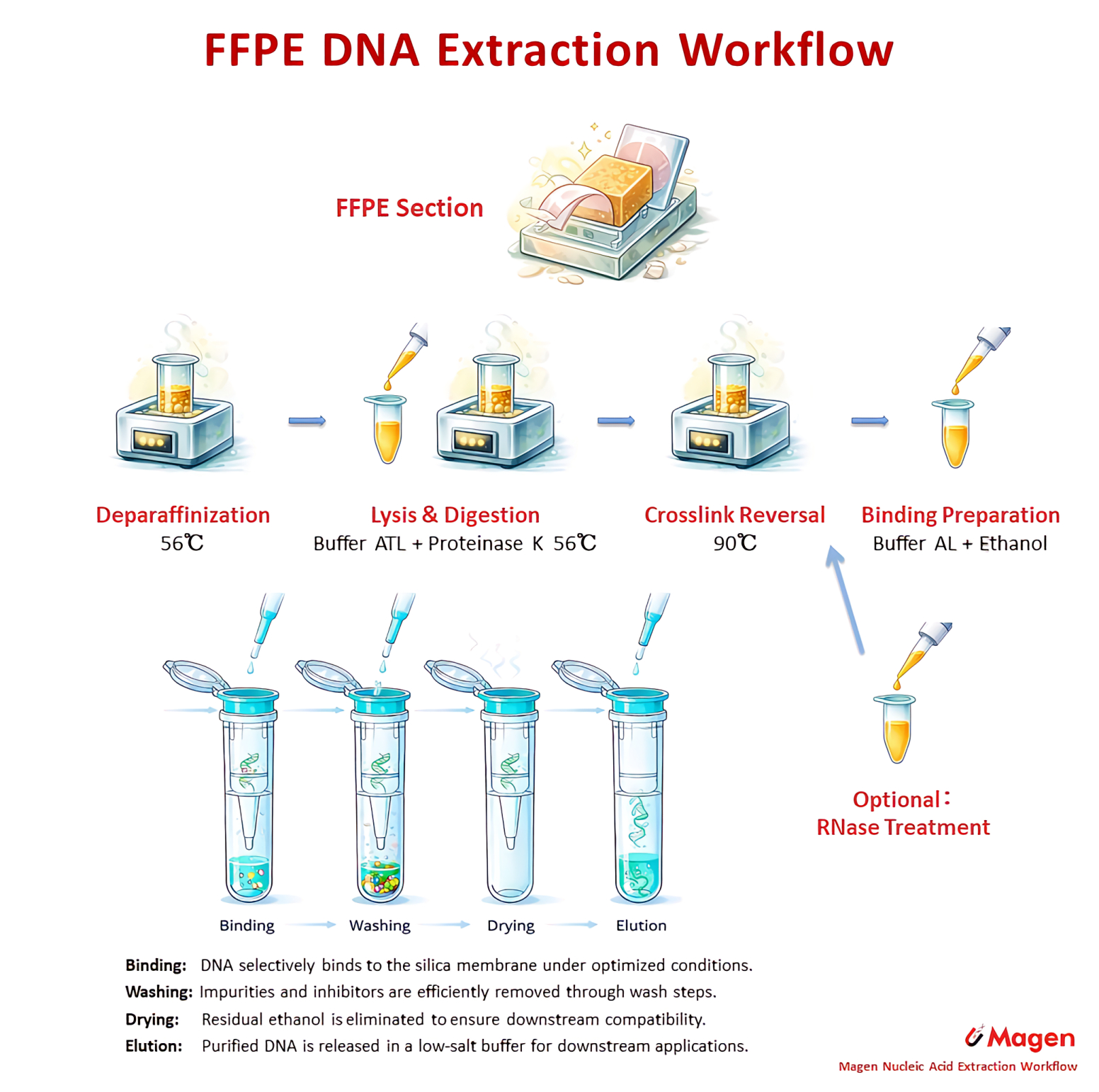
Figure 1. Representative FFPE DNA column workflow routes
The fragment-selection magnetic DNA route is represented by D6323B MagPure FFPE DNA Kit. This route is not simply a magnetic version of routine DNA extraction. Its defining feature is the binding-condition adjustment stage, where Buffer BD-controlled binding conditions influence short-fragment recovery behavior. This makes the route relevant for NGS-oriented FFPE DNA workflows, where fragment profile can affect downstream library preparation and mutation analysis.
The high-purity magnetic DNA route is represented by D6323D MagPure FFPE DNA Kit (High Pure). This route was developed around binding strategy rather than fragment selection. High Salt Bind and Ethanol Bind are treated as alternative binding methods, allowing the workflow to address different purity and recovery pressures. In this route, the key question is not only how much DNA can be recovered, but how well co-purified material and assay-interfering carryover can be controlled.
2.2 RNA-Only Workflow Routes
FFPE RNA workflows are not shortened DNA protocols. They are built around a different balance: sufficient tissue digestion and RNA release must be achieved without unnecessary heat exposure or downstream DNA interference. Because FFPE RNA is often fragmented before extraction begins, the workflow must avoid adding further stress while still reversing enough fixation-related modification to support downstream analysis.
The column RNA route is represented by R4143 HiPure FFPE RNA Kit. This workflow uses controlled digestion and heat treatment before silica membrane RNA purification. R4144 HiPure FFPE RNA Plus Kit extends this route by integrating DNase treatment, which becomes important when downstream assays require RNA-derived signal to be interpreted with reduced genomic DNA contribution.
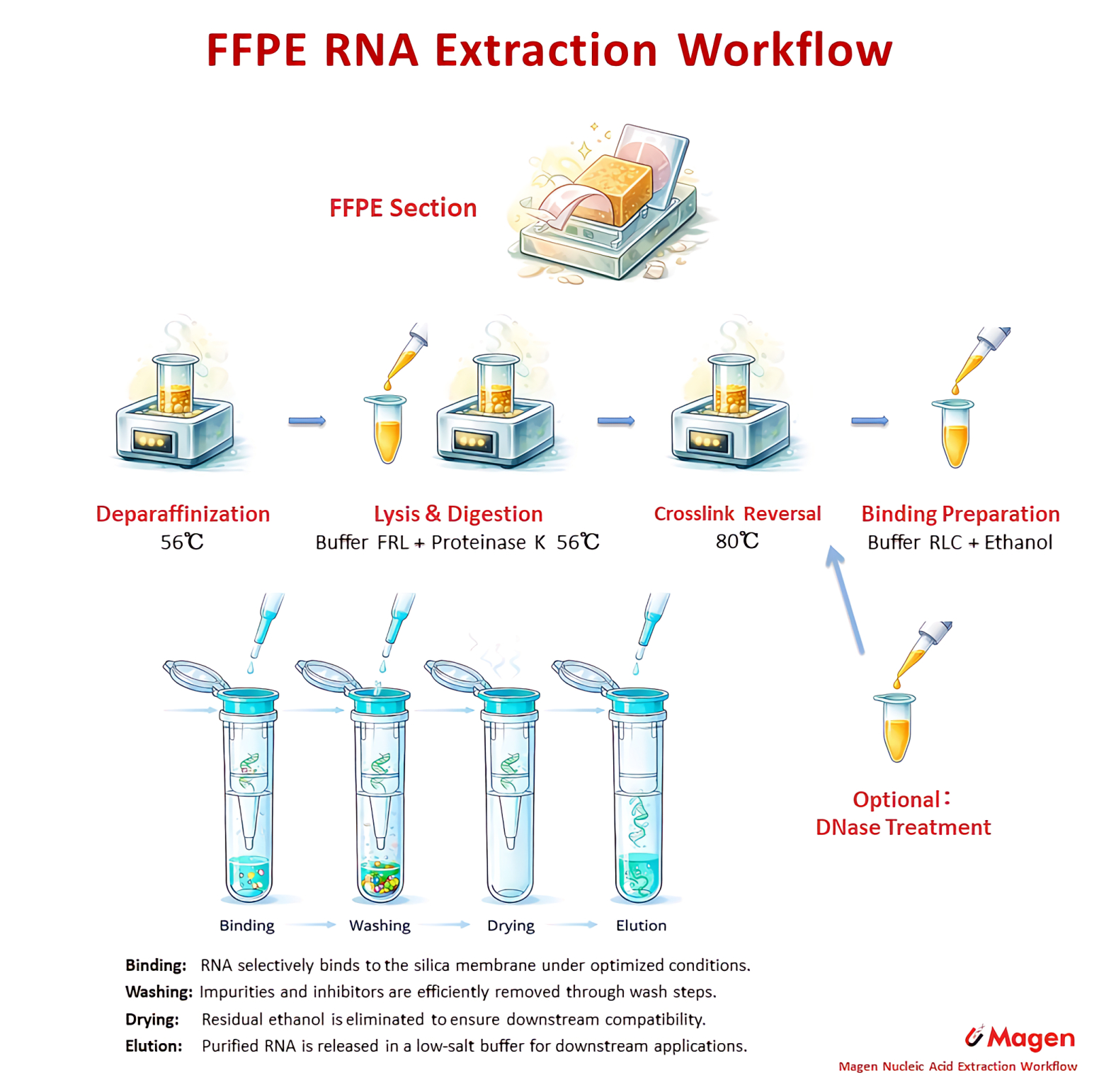
Figure 2. Representative FFPE RNA column workflow routes, showing controlled heat treatment and DNase-integrated RNA preparation.
The magnetic RNA route is represented by IVD3022 MagPure FFPE RNA Kit. This workflow uses magnetic particles for RNA binding and includes DNase I treatment as part of the magnetic purification route. Compared with the column RNA route, the downstream logic is based on magnetic binding, magnetic separation, wash control, DNase treatment and elution rather than membrane loading.
Together, these RNA routes reflect a central design principle: FFPE RNA extraction is not only about recovery. It is also about controlled heat exposure, residual DNA management and downstream RT-PCR or gene expression compatibility.
2.3 DNA/RNA Co-Extraction Workflow Routes
DNA/RNA co-extraction is not defined only by the fact that both analytes are recovered from the same FFPE input. The more important question is where and how DNA and RNA are separated within the workflow. In the Magen FFPE system, the co-extraction routes can be understood through two main strategies: early partition and sequential adsorption.
Early Partition Strategy
The early partition strategy is represented by IVD5116 HiPure FFPE DNA/RNA Kit and IVD3026 MagPure FFPE DNA/RNA Kit. In both workflows, FFPE sections first enter a shared pretreatment route. After deparaffinization, protease-assisted lysis and centrifugation, the workflow separates the RNA-containing supernatant from the DNA-containing pellet. From this point onward, the two nucleic acid fractions are processed through separate purification paths.
The difference between IVD5116 and IVD3026 is not the separation principle, but the downstream purification format. IVD5116 uses column purification after early partition, while IVD3026 uses magnetic bead purification after early partition. This means that both workflows share the same core partitioning logic, but differ in how the separated DNA and RNA fractions are captured, washed and eluted.
The technical value of this strategy is that the RNA fraction can be removed from the digestion residue and DNA-oriented processing path at an early stage. This allows the RNA-containing supernatant to enter a more RNA-oriented route, while the DNA-containing pellet can undergo additional lysis and high-temperature treatment before DNA purification. For this reason, early partition is particularly useful when preserving RNA quality or RNA integrity is a major workflow concern.
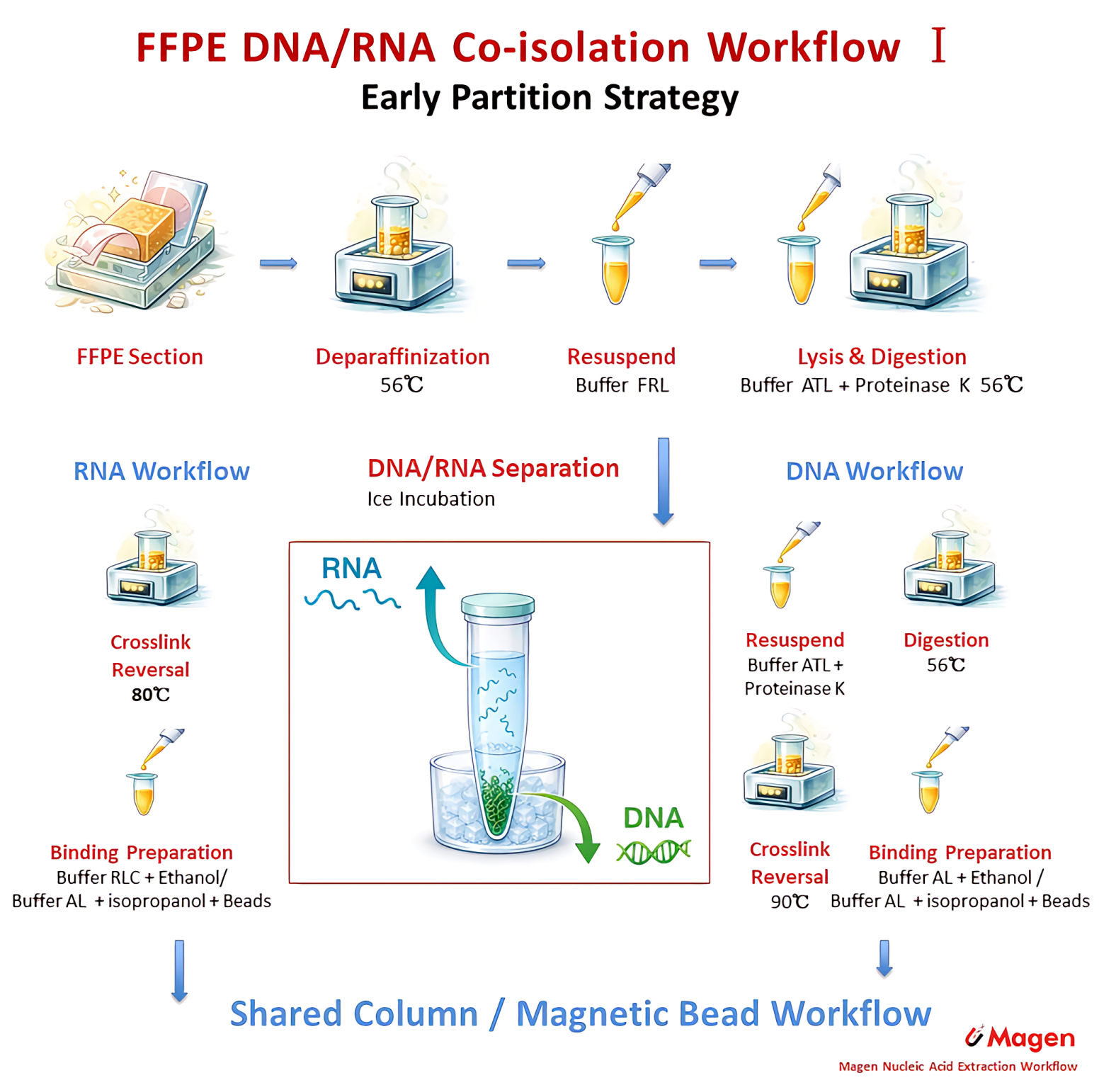
Figure 3A. Early partition strategy for FFPE DNA/RNA co-extraction, showing shared FFPE pretreatment followed by separation of the RNA-containing supernatant and DNA-containing pellet.
Sequential Adsorption Strategy
The sequential adsorption strategy is represented by R6327 MagPure FFPE DNA/RNA Kit. In this workflow, DNA and RNA are not separated immediately as pellet and supernatant after the early pretreatment stage. Instead, the FFPE lysate first enters a magnetic adsorption step in which DNA is captured by magnetic particles. The RNA-containing supernatant is then transferred into a second magnetic binding route for RNA purification.
This makes the first DNA-binding step the central partition point of the workflow. In contrast to the early partition strategy, where physical separation of the DNA-containing pellet and RNA-containing supernatant defines the split, sequential adsorption separates the two analytes through staged magnetic binding behavior.
The technical value of sequential adsorption is that DNA and RNA are first released within a shared lysate environment before being recovered through staged magnetic binding. This route is therefore oriented toward high-recovery DNA/RNA co-extraction, especially when the workflow objective is to obtain both analytes from the same FFPE input for parallel analysis or multi-omics-oriented applications. Compared with early partition, its emphasis is less on early RNA fraction isolation and more on efficient sequential recovery of both DNA and RNA.
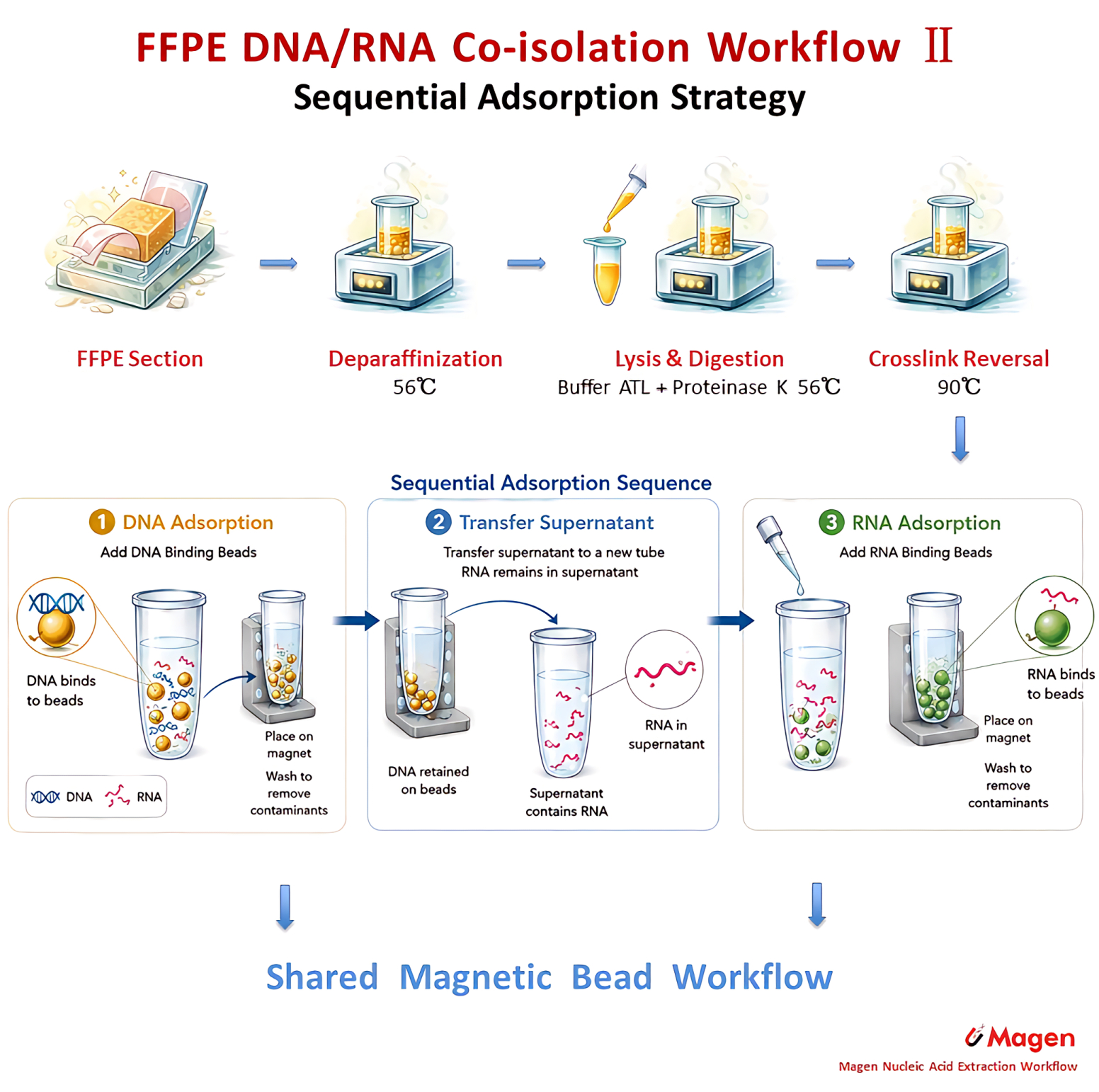
Figure 3B. Sequential adsorption strategy for FFPE DNA/RNA co-extraction, showing DNA-first magnetic capture followed by RNA purification from the remaining supernatant.
In simple terms, early partition prioritizes early RNA-oriented fraction handling, while sequential adsorption prioritizes staged recovery of both DNA and RNA from a shared lysate system.
3.0 Development Logic Behind the Workflow Routes
The FFPE workflow system did not arise from one universal extraction logic applied to every sample. It took shape because each analyte and each downstream use places different pressure on the process. The following development observations are useful not because they form a large performance study, but because they show how small process choices became route-defining decisions.
3.1 Heat Treatment and Fraction Handling in Early Partition Co-Extraction
In early partition co-extraction workflows, the DNA and RNA branches require different handling priorities. Process testing with DNA marker input showed higher recovery under 90℃ treatment than under 70℃ treatment, supporting the stronger high-temperature reversal step used in the DNA branch. In contrast, RNA process testing showed better Poly A recovery when the RNA-containing fraction was kept on ice rather than held at room temperature. The same FFPE input therefore enters a workflow in which the DNA branch and RNA branch are deliberately managed differently.
3.2 Controlled Heat Exposure in RNA Workflows
The RNA column route was evaluated from a related but separate angle. Under the tested RNA extraction conditions, controlled 80℃ treatment did not produce visible RNA degradation by gel analysis. In RT-PCR-oriented validation, viral RNA introduced into lysis buffer produced Ct values close to reference material after extraction, without obvious amplification inhibition in that test setting.
Archived FFPE material gives this route practical relevance. FFPE blocks may preserve clinically annotated tissue for retrospective molecular studies, but fixation, embedding and storage can introduce fragmentation, chemical modification and protein-nucleic acid crosslinking. In FFPE RNA route testing, one-year archived FFPE animal tissues, including liver, muscle and heart-derived samples, still generated measurable RNA after extraction. This supported the practical relevance of the column RNA route for archived FFPE materials. DNase-related evaluation further clarified the role of DNA carryover control, especially when downstream qRT-PCR results need to be interpreted as RNA-derived signal rather than residual genomic DNA signal.
3.3 Magnetic DNA Route Development
The DNA-oriented magnetic routes developed around different observations. In the fragment-selection route, Buffer BD-controlled binding conditions were evaluated for their effect on short-fragment recovery behavior. In the high-purity route, gel-well retention and co-purified material became practical indicators that binding strategy mattered, not only total recovery. These observations guided the separation between a fragment-selection route and a high-purity route.
3.4 Small Process Variables That Define Route Behavior
A temperature setting, an ice-hold step, the position of DNase digestion, the particle binding chemistry or the volume of a binding buffer may look secondary in a protocol. In practice, these details define whether a route is oriented toward recovery, preservation, partitioning, fragment behavior or downstream compatibility.
| Development Observation | Technical Meaning for Route Design |
|---|---|
| 90℃ treatment in the DNA branch | Supports stronger reversal conditions when DNA recovery is the main pressure point. |
| Ice handling of the RNA-containing fraction | Reflects the need to protect RNA handling conditions during co-extraction. |
| Controlled 80℃ RNA treatment | Supports heat-assisted RNA recovery when exposure is controlled rather than extended. |
| Buffer BD-dependent short-fragment behavior | Turns magnetic binding into a fragment-selection control point. |
| Gel-well retention in some magnetic DNA routes | Highlights why high-purity binding chemistry can matter beyond yield. |
| DNase-integrated RNA routes | Addresses residual DNA as a downstream interpretation issue, especially for qRT-PCR-oriented work. |
4.0 Workflow Time, Operational Complexity and Detailed Notes
Processing time differs because the routes place time in different parts of the workflow: digestion, heat treatment, partitioning, DNase treatment, magnetic binding, bead drying or elution. Co-extraction workflows require particular care in interpretation. A dual-output workflow is complete only after both analyte paths have been processed.
| Route Type | Representative Workflow | Typical Processing Time | Time Interpretation |
|---|---|---|---|
| Routine DNA column route | D3126 | Approx. 2.5-14 h | Longer range reflects sample-dependent digestion. |
| RNA column route | R4143 / R4144 | Approx. 1.5-2 h | Shorter RNA route with DNase branch where required. |
| Early Partition Column Co-Extraction | IVD5116 | Approx. 4.5 h | Shared pretreatment plus both RNA and DNA column paths. |
| Early Partition Magnetic Co-Extraction | IVD3026 | Approx. 4.7 h | Shared pretreatment plus both magnetic paths. |
| Sequential Adsorption Magnetic Co-Extraction | R6327 | Approx. 3.3-14.5 h | Range reflects sample-dependent digestion before sequential binding. |
| Fragment-selection DNA route | D6323B | Approx. 3 h | Binding-condition adjustment adds route-specific control. |
| Dual-binding high-purity DNA route | D6323D | Approx. 2.9 h per method | High Salt Bind and Ethanol Bind are alternative methods. |
| Magnetic RNA route with DNase | IVD3022 | Approx. 1.8 h | Magnetic RNA route with integrated DNase treatment. |
Time is not only a convenience metric. It reveals where a route places its technical burden. A DNA route may spend most of its time in digestion and reversal. A magnetic route may add repeated separation and drying steps. A co-extraction route adds handling complexity because two fractions must be preserved and processed rather than collapsed into one purified output.
4.1 Detailed Workflow Notes
The time estimates above provide a structural comparison of the routes. For step-by-step handling logic, estimated processing time and route-specific interpretation, the detailed workflow notes provide a more practical view of each route.
| Workflow Route | Detailed Workflow Note |
|---|---|
| FFPE DNA column route | D3126 HiPure FFPE DNA Kit Workflow Note |
| FFPE RNA column route | R4143 / R4144 HiPure FFPE RNA Workflow Note |
| Early Partition Column Co-Extraction | IVD5116 HiPure FFPE DNA/RNA Workflow Note |
| Early Partition Magnetic Co-Extraction | IVD3026 MagPure FFPE DNA/RNA Workflow Note |
| Sequential Adsorption Magnetic Co-Extraction | R6327 MagPure FFPE DNA/RNA Workflow Note |
| Fragment-selection FFPE DNA route | D6323B MagPure FFPE DNA Workflow Note |
| Dual-binding high-purity FFPE DNA route | D6323D MagPure FFPE DNA Workflow Note |
| Magnetic FFPE RNA with DNase | IVD3022 MagPure FFPE RNA Workflow Note |
5.0 Pre-Analytical and Workflow Considerations
FFPE workflow performance should always be interpreted together with sample history. Fixation time, block age, section thickness, tissue density, storage condition and carryover material can all influence what is ultimately recovered. The same workflow route may behave differently when applied to surgical tissue, biopsy-derived material or wax roll input.
Deparaffinization prepares the sample for lysis, but it does not determine the full outcome by itself. Digestion completeness, phase or pellet handling, magnetic separation quality and drying control often have stronger influence on recovery stability and downstream compatibility. In magnetic workflows, abnormal gel-well behavior may indicate co-purified material or incompletely removed particles, and should be interpreted together with concentration and downstream assay results.
OD ratios also require context. In FFPE-derived DNA and RNA workflows, A260/280, A260/230, Qubit concentration, gel appearance and amplification performance may not always tell the same story. For example, RNA datasets may show A260/280 values close to the expected range while A260/230 varies across samples. This is why FFPE workflow evaluation should not be reduced to a single yield or purity number.
6.0 Downstream Application Orientation
Downstream applications create different pressure on the extraction route. Routine PCR or mutation detection depends mainly on amplifiable FFPE DNA. qPCR and inhibitor-sensitive assays may require closer attention to carryover material, purity and binding chemistry. NGS-oriented workflows may be affected by fragment profile and short-fragment behavior. RT-PCR and gene expression workflows place greater emphasis on RNA preservation, controlled heat exposure and DNA carryover control. Parallel DNA/RNA analysis adds another layer of complexity, because both fractions must be preserved from the same FFPE input.
| Downstream Pressure | Workflow Implication |
|---|---|
| Routine PCR / mutation detection | Requires amplifiable FFPE DNA through a stable DNA recovery route. |
| qPCR or inhibitor-sensitive assays | Purity, co-purified material and binding strategy may matter as much as apparent yield. |
| NGS / mutation profiling | Fragment profile and short-fragment behavior can influence library preparation and target representation. |
| RT-PCR / gene expression | RNA recovery, controlled heat exposure and DNA carryover control become central. |
| DNA-sensitive RNA assays | DNase-integrated RNA workflows help reduce the risk that residual genomic DNA affects result interpretation. |
| Parallel DNA/RNA analysis | The route must preserve and process both analyte fractions from the same FFPE input. |
| Multi-omics-oriented workflows | Early partition or sequential adsorption routes can align DNA and RNA recovery within one sample-processing system, depending on whether the priority is RNA-oriented handling or high-recovery co-extraction. |
These mappings should be read as workflow tendencies rather than rigid boundaries, because FFPE sample condition and assay sensitivity remain decisive variables.
7.0 Closing Note
Taken together, these routes define a practical FFPE workflow system rather than a single extraction format with minor variations. DNA, RNA and DNA/RNA co-extraction each place different pressure on the sample-processing steps. Fragment selection, high-purity binding and DNase treatment add further route-specific requirements.
Understanding where the routes diverge — at lysis, heat treatment, partitioning, binding, washing or elution — makes it easier to relate each workflow to the real constraints of archived FFPE material and the requirements of downstream molecular analysis.


