Introduction
MagPure Circulating DNA Maxi Kit (IVD5435) represents the magnetic bead configuration within the Magen circulating DNA portfolio. Developed through long-term refinement of magnetic particle chemistry, the system is designed for scalable plasma input ranging from 1 to 8mL.
Rather than focusing solely on yield claims, development emphasized bead surface consistency, adsorption stability and workflow adaptability. The extraction chemistry supports both manual magnetic stands and automation-oriented liquid handling systems.
In internal validation, plasma extraction results showed reproducible cfDNA yield across replicates. Comparative evaluations under equivalent conditions indicated performance consistent with established European magnetic systems. Stability testing across production batches demonstrated no measurable drift in physicochemical parameters over extended storage periods.
Laboratories preferring membrane-based workflows may refer to (HiPure Circulating DNA Kit – IVD3182). For lower plasma volumes or fragment-selective workflows, see:
· Small-volume magnetic extraction → (IVD5432)
· Size-selective enrichment → (12927)
· Column-based micro plasma extraction → (D3180)
IVD5435 serves as the scalable magnetic reference model within the system.
Details
Workflow
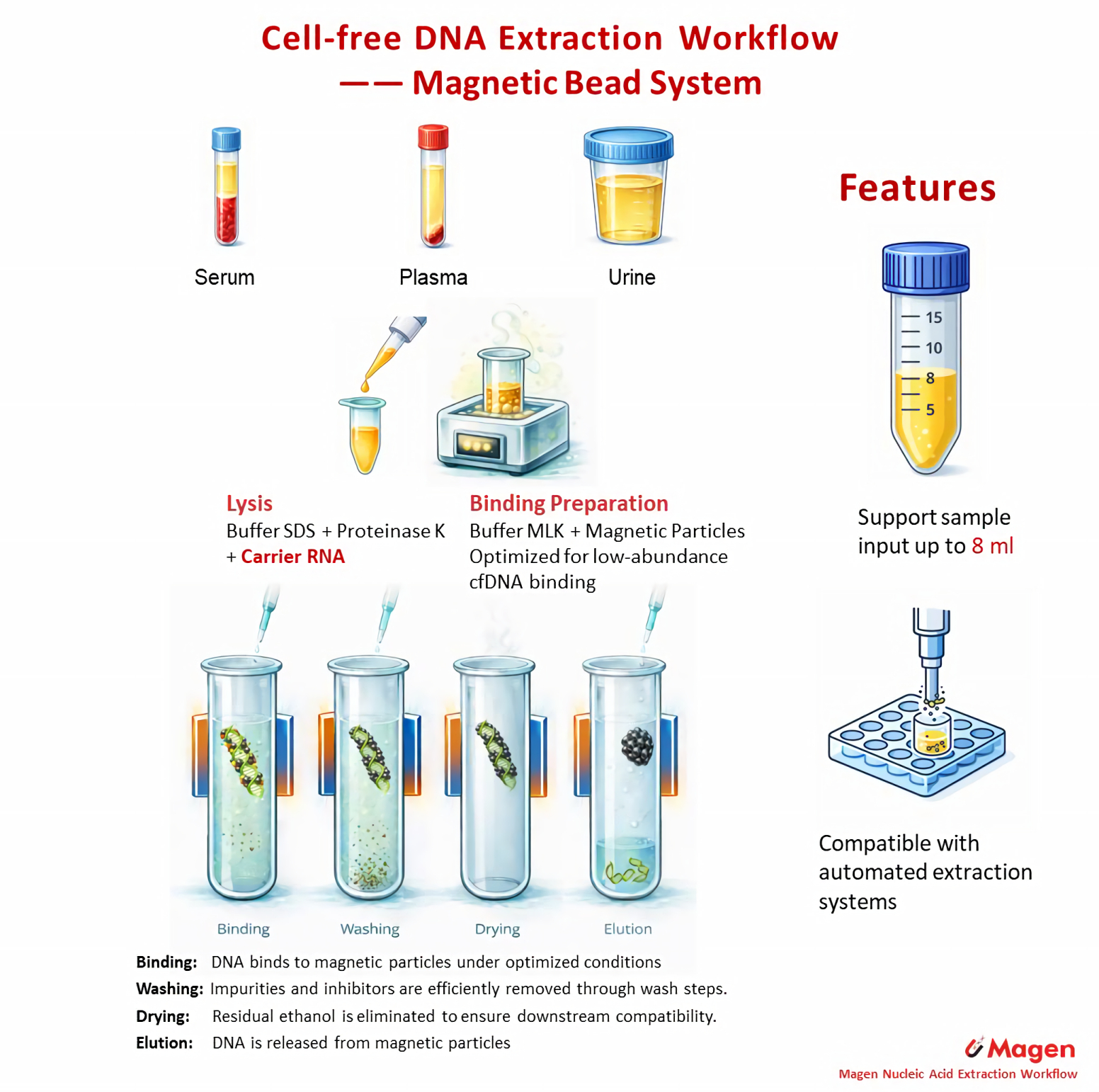
Workflow Overview
The MagPure Circulating DNA Maxi Kit follows a magnetic bead–based workflow for purification of circulating DNA from plasma and serum. DNA is captured through particle–DNA interaction under mixing conditions rather than membrane flow-through, enabling flexible handling of larger sample volumes and low-abundance cfDNA.
Sample Handling Logic
This workflow supports 1–8 mL plasma or serum input. For lower input volumes, DNA binding is typically completed in a single step, while larger-volume samples are processed through staged binding to maintain recovery consistency. Magnetic separation replaces column flow-through, allowing scalable handling and compatibility with automated systems.
Time and Workflow Characteristics
Compared to column workflows, processing time is typically longer due to mixing, magnetic separation and repeated liquid handling steps. Under standard conditions, the workflow is generally completed within about 105–130 minutes. For detailed workflow steps, binding strategies and processing guidance, please refer to the Workflow Note in the Download section.
Engineering Characteristics
Magnetic Particle Consistency
Magnetic bead solid content and responsiveness are monitored during production. Experimental evaluation demonstrated that bead concentration directly influences recovery efficiency, and optimized bead-to-volume ratios are applied in the final workflow.
Plasma Recovery Stability
Across 4–8mL plasma processing, replicate extractions maintained stable Qubit concentration with minimal deviation, supporting higher-volume workflows.
Storage and Lot Stability
Physicochemical parameters (pH, density, conductivity) and marker recovery testing confirmed consistent performance across multiple production batches, including kits stored up to two years.
Workflow Scalability
IVD5435 integrates into a broader extraction system:
Technical Validation
Marker Recovery Testing
Spike-in marker experiments demonstrated DNA recovery efficiencies typically above 80% across multiple reagent batches, confirming stable adsorption performance of the magnetic particle system.
Plasma cfDNA Extraction Performance
Extraction experiments using plasma samples showed circulating DNA yields typically around 5–7 ng per mL plasma when processing milliliter-scale plasma volumes.
Comparative Extraction Evaluation
Under equivalent extraction conditions, circulating DNA yields obtained using the MagPure system were comparable to those produced by widely used column-based circulating DNA extraction kits.
Application Scenario Summary
The following table summarizes selected published research scenarios in which IVD5435 or closely related MagPure circulating DNA formats were used for cfDNA preparation. The purpose is to help users understand typical application directions of this magnetic bead-based product route before reviewing the related technical notes and publications.
|
Application Scenario
|
Related Format
|
Sample / cfDNA Source
|
Downstream Research Use
|
|
Maternal plasma cfDNA-based NIPT for hemoglobin Bart hydrops fetalis risk in α-thalassemia carrier couples
|
IVD5435
|
Maternal plasma cfDNA / cffDNA from pregnancies in which both parents carried the α-thalassemia SEA deletion
|
Target-captured sequencing with UID adapters, fetal fraction estimation and Bayesian genotype classification for fetal SEA deletion status, hemoglobin Bart hydrops fetalis detection and concurrent T13 / T18 / T21 screening
|
|
Maternal plasma cfDNA-based noninvasive prenatal diagnosis for TSC single-gene disorder risk
|
IVD5432
|
Maternal plasma cfDNA and matched maternal wbcDNA from pregnancies at risk of TSC
|
Targeted NGS of maternal plasma cfDNA and wbcDNA for fetal TSC1 / TSC2 genotype deduction and NIPD feasibility evaluation
|
View more application scenarios
|
Application Scenario
|
Related Format
|
Sample / cfDNA Source
|
Downstream Research Use
|
|
Maternal plasma cfDNA fragmentomics for gestational diabetes mellitus prediction and molecular characterization
|
IVD5432
|
Longitudinal maternal plasma cfDNA from women with GDM and matched healthy pregnant controls
|
High-depth cfDNA sequencing and integrative fragmentomics / TSS-score analysis for lipid-metabolism association, PRSS1-related pancreatic marker evaluation and early GDM prediction modeling
|
|
CRC plasma cfDNA methylation assay development for colorectal cancer and precancerous lesion detection
|
IVD5435
|
Plasma cfDNA from CRC, precancerous lesion and no-evidence-of-disease participants
|
Multiplex TaqMan real-time PCR for SEPT9 / SDC2 / ALX4 methylation detection and CRC / precancerous lesion screening performance evaluation
|
|
Breast cancer plasma cfDNA methylation biomarker discovery for early non-invasive diagnosis research
|
IVD5435
|
Plasma cfDNA from early-stage breast cancer patients and healthy female controls
|
Target-capture bisulfite sequencing and BSP-HiSeq validation for breast cancer-associated cfDNA CpG methylation biomarker discovery and diagnostic performance evaluation
|
|
Echinococcus granulosus cfDNA monitoring before and after albendazole treatment in cystic echinococcosis
|
IVD5432
|
Plasma Eg cfDNA from cystic echinococcosis patients before and after albendazole treatment
|
High-throughput sequencing for Eg cfDNA read identification, fragment-length analysis, genomic release-source comparison and albendazole treatment-response monitoring research
|
Note: These examples describe selected published research uses of IVD5435 or closely related MagPure circulating DNA formats for upstream cfDNA preparation. The extraction workflow provides purified nucleic acid input for downstream assays; final research conclusions depend on the complete study design, sequencing or detection method, bioinformatics analysis and clinical validation strategy.
Kit Contents
|
Contents
|
IVD5435
|
|
Purification Times
|
50
|
|
MagPure Particles F
|
14 ml
|
|
Carrier RNA
|
310 μg
|
Proteinase K
|
240 mg
|
Protease Dissolve Buffer
|
15 ml
|
Buffer SDS
|
15 ml
|
|
Buffer MLK
|
500 ml
|
|
Buffer MAW1
|
250 ml
|
|
Buffer MW2*
|
50 ml
|
|
Elution Buffer
|
60 ml
|
Storage and Stability
Proteinase K, Carrier RNA and MagPure Particles F should be stored at 2–8°C upon arrival. However, short-term storage (up to 12 weeks) at room temperature (15–25°C) does not affect their performance.Theremaining kit components can be stored dry at room temperature (15–25°C) and are stable for at least 18 months under these conditions.The entire kit can be stored at 2–8°C, but in this case buffers should beredissolved before use. Make sure that all buffers are at room temperature when used.
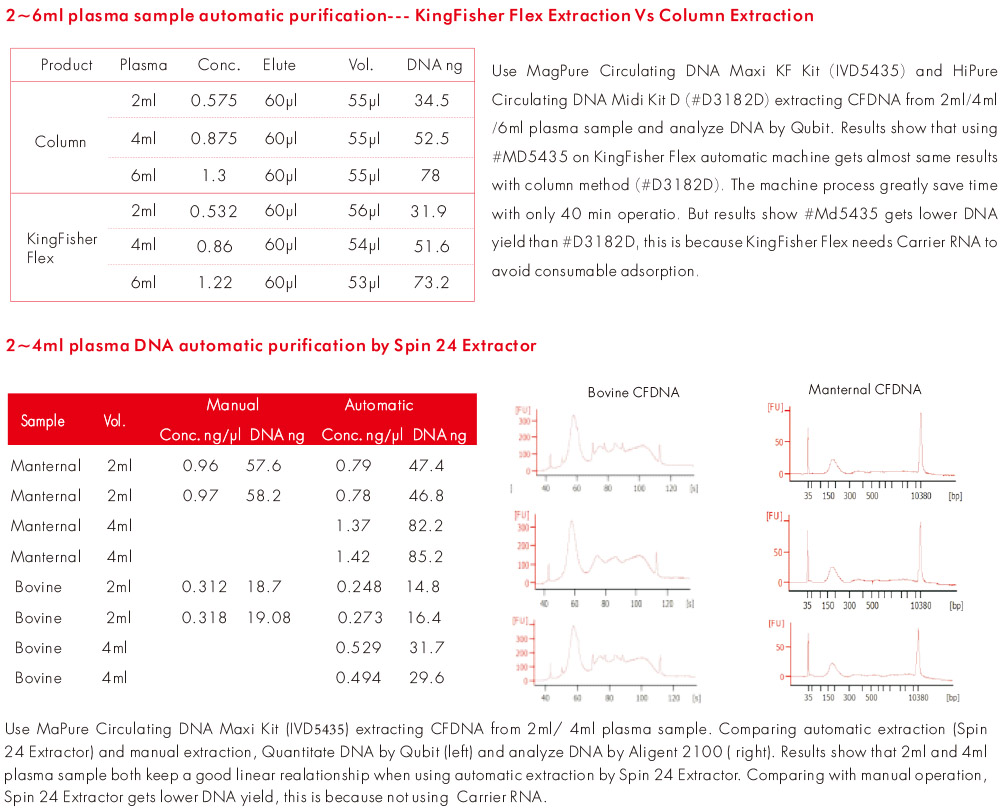
Experiment Data
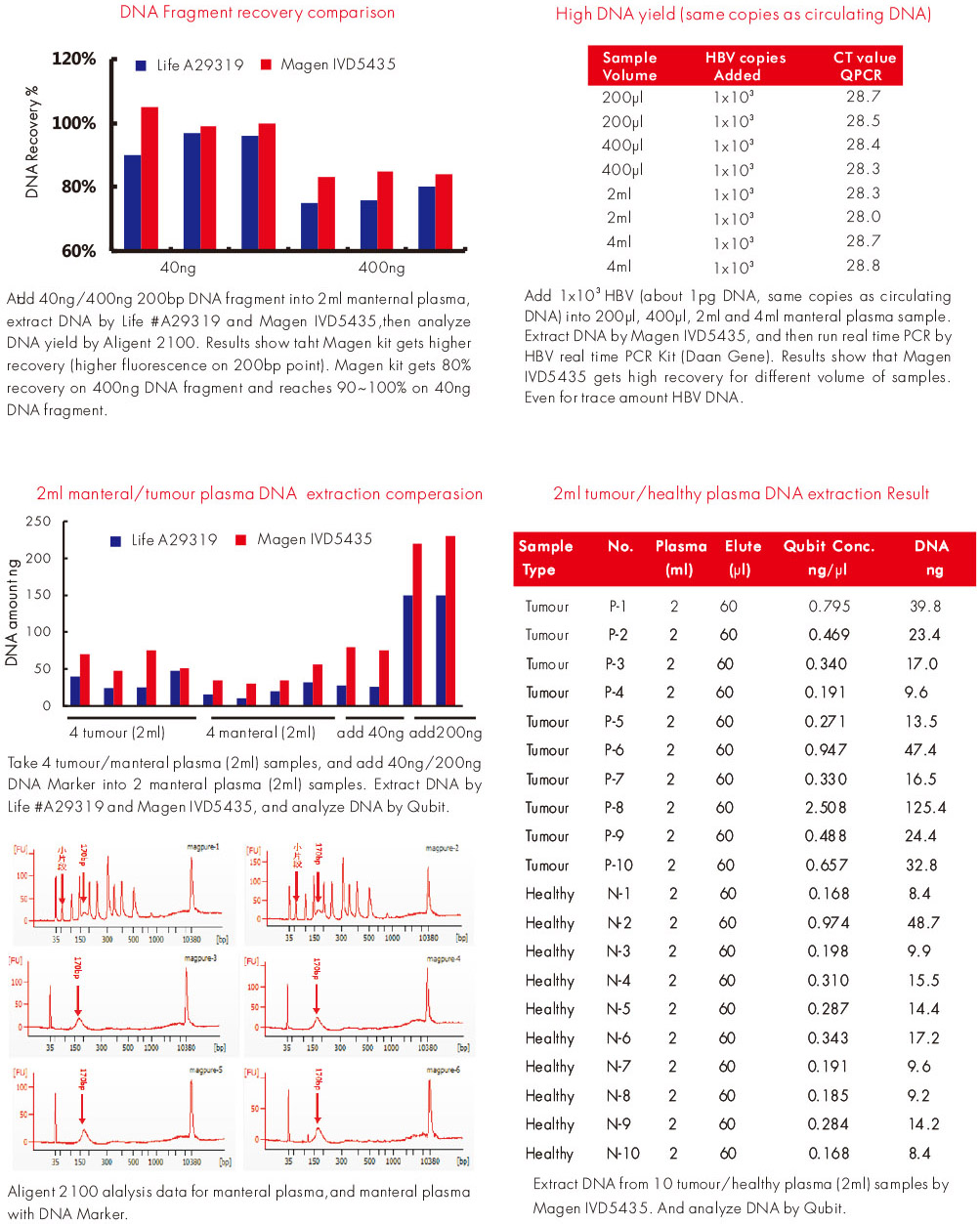
Purchase Guide
For guidance on selecting the most appropriate DNA extraction system based on sample type, input volume and workflow requirements:
👉 Magen Kits Selection Guide
For a broader technical overview of cfDNA workflow routes, processing logic and downstream application orientation:
👉 cfDNA Extraction & Enrichment Workflows
For reference workflow structure and comparative processing logic across representative cfDNA workflows:
👉 cfDNA Workflow Notes (IVD3182 / IVD5435 / 12927)
Related cfDNA Application Notes: These application notes help explain how this circulating DNA extraction route fits different plasma cfDNA research workflows and downstream assay requirements.
👉 Application Map · Methylation Biomarkers · Prenatal cfDNA